
Spring/Summer 1999
Volume 6, Issue 2
Spring/Summer 1998
Volume
3, Issue 1
January 1995
Volume
2, Issue 4
October 1994
Volume
2, Issue 1
January 1994
RESEARCH FOCUS: EDUCATION
The W. M. Keck Center Expands its Training and Research in Computational Biology
The W. M. Keck Center for Computational Biology, a joint effort of Rice University, Baylor College of Medicine, and the University of Houston, was established originally by the W .M. Keck Foundation for the purpose of fostering collaboration among biologists, biomedical researchers, computer scientists and mathematicians to develop and deploy new, more powerful tools for biological research. Five years later, the Center has matured into a self-supporting operation that attracts a broad base of funding for its innovative training programs that integrate the computational aspects of simulation, imaging, and sequence analysis with a solid foundation in biophysics, biochemistry and genetics.
Three of the Keck Center's major supporters are the Dunn Foundation, the National Library of Medicine (NLM), and the National Science Foundation (NSF). A year ago, the NSF awarded the center $1.5 million over five years in support of the cutting-edge computational science training programs that Keck offers to high school through postdoctoral students. As a result, the center is fueling its successful graduate study program and is expanding its training efforts with high school, undergraduate, and postdoctoral trainees by paying for tuition, stipends, and some research expenses.
Participants in the training programs are drawn from various disciplines that include computer science, engineering, biology, biochemistry, chemistry, mathematics, neuroscience, physics, and biophysics. Graduate students are admitted to programs at either Baylor, Rice, or University of Houston, and once accepted, may apply to become a fellow in the Keck Center. Keck Fellows have access to the advanced computational and analytical resources of the center.
Postdoctoral scientists are chosen for their demonstrated abilities in research of interest to the Keck Center faculty, focused mainly in the areas of computational crystallography, computational imaging, genome informatics, computational biochemistry, and advanced computational methodologies. The trainees participate in seminars, symposia, and other research-related activities. In keeping with the interdisciplinary spirit of the program, all postdoctoral scientists have two mentors, one in biological science and one in computational science.
A new NSF-supported training program debuted this summer, in which 22 undergraduate students and a few high school students spent two to three months working on intensive research projects in computational biology. Their mentors were Keck Center professors, scientists, and visiting researchers from other related departments at Rice, Baylor, and the University of Houston. This program will continue during the academic year for 12 to 15 students of high academic standing and of diverse backgrounds.
The Keck Center will also continue its popular weekly seminar program, established early in the life of the Center. The seminars, which feature Keck faculty, postdoctorate, and graduate speakers as well as guest experts in computational science and biology, are held every Friday from 4:00 to 5:30 p.m. at Rice and are open to anyone who is interested. Past subjects have included "Space efficient multisequence alignment," "Structural studies of actin binding proteins," "Languages for parallel computation," and "Molecular dynamics simulations of peptides, proteins, and nucleic acids." The seminars are advertised by email. To get on the email list, contact georgep@rice.edu .
The success and growth of all training and research programs at the Keck Center are based on the strong collaboration of its members, and its close ties with other centers and scientists that work in related fields. The CRPC, along with the Texas Center for Advanced Molecular Computation (TCAMC), has been closely involved with the Keck Center, providing expertise and resources in the area of advanced high- performance computing. Other organizations that work with the Keck Center include the Human Genome Center, the 3-Dimensional Electron Microscopy Resource Center, and the Molecular Biology Computational Research Center at Baylor, and the Institute for Biosicences and Bioengineering at Rice.
"Collaboration has made us the successful organization we are today," says Rice Keck Center Training Director George Phillips, who is also Professor of Biochemistry and Cell Biology. "We complement one another's strengths and come out with a whole that is more than the sum of its parts. We are viewed as a real community of scholars, which benefits the students and drives the success of our research projects and funding opportunities."
"Our goal is always to be inclusive rather than exclusive," says Keck Executive Director Marc Archambault of the collaboration efforts. "We want to define the emerging field of computational science as broadly as possible and make use of research and ideas from a wide range of fields that can contribute to and benefit from this groundbreaking discipline."
For more information, contact compbio@rice.edu , the URL http://www.rice.edu , or write to the W. M. Keck Center for Computational Biology, MS 141, Rice University, 6100 Main Street, Houston, TX 77005.
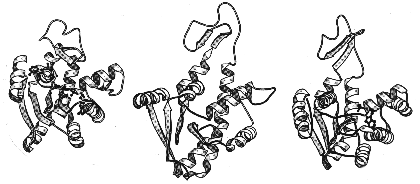
Closed Simulation Open
The enzyme adenylate kinase has been studied by both experiment and computer simulation. The enzyme has a lid (shown at top) that opens and closes as a part of its bilogical action. This opening has been simulated using computers that solve Newton's equations of motion for each of the two thousand atoms in the enzymei. This is but one example of many novel uses of computers in biological research at the Keck Center.
Table of Contents