 Proteins are not rigid static structures, but exist as a complex
dynamical ensemble of closely related substates. It
is the dynamics of the system that permit transitions between these
conformational substates and there is evidence that this
dynamic behavior is also responsible for the kinetics of ligand entry
and ligand binding in systems such as myoglobin.
Proteins are not rigid static structures, but exist as a complex
dynamical ensemble of closely related substates. It
is the dynamics of the system that permit transitions between these
conformational substates and there is evidence that this
dynamic behavior is also responsible for the kinetics of ligand entry
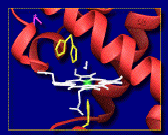
and ligand binding in systems such as myoglobin. X-ray crystallography is able to provide high resolution maps of the average structure of a protein, but this structure only represents a static conformation. Molecular dynamics, which solves the Newtonian equations of motion for each atom , is not subject to such approximations and can be used to model the range of motions available to the protein on short (nanosecond) time-scales.
If the ensemble of conformations is thought of as a matrix , then anayltical tools from linear algebra can be used. Here the tool of choice is the SVD . The SVD can be used for both compressing the data required to represent the simulation and for locating interesting functional behavior.